Binomial confidence intervals using Bayesian inference
binom.bayes.RdUses a beta prior on the probability of success for a binomial distribution, determines a two-sided confidence interval from a beta posterior. A plotting function is also provided to show the probability regions defined by each confidence interval.
binom.bayes(x, n,
conf.level = 0.95,
type = c("highest", "central"),
prior.shape1 = 0.5,
prior.shape2 = 0.5,
tol = .Machine$double.eps^0.5,
maxit = 1000, ...)
binom.bayes.densityplot(bayes,
npoints = 500,
fill.central = "lightgray",
fill.lower = "steelblue",
fill.upper = fill.lower,
alpha = 0.8, ...)Arguments
- x
Vector of number of successes in the binomial experiment.
- n
Vector of number of independent trials in the binomial experiment.
- conf.level
The level of confidence to be used in the confidence interval.
- type
The type of confidence interval (see Details).
- prior.shape1
The value of the first shape parameter to be used in the prior beta.
- prior.shape2
The value of the second shape parameter to be used in the prior beta.
- tol
A tolerance to be used in determining the highest probability density interval.
- maxit
Maximum number of iterations to be used in determining the highest probability interval.
- bayes
The output
data.framefrombinom.bayes.- npoints
The number of points to use to draw the density curves. Higher numbers give smoother densities.
- fill.central
The color for the central density.
- fill.lower,fill.upper
The color(s) for the upper and lower density.
- alpha
The alpha value for controlling transparency.
- ...
Ignored.
Details
Using the conjugate beta prior on the distribution of p (the probability of success) in a binomial experiment, constructs a confidence interval from the beta posterior. From Bayes theorem the posterior distribution of p given the data x is:
p|x ~ Beta(x + prior.shape1, n - x + prior.shape2)
The default prior is Jeffrey's prior which is a Beta(0.5, 0.5)
distribution. Thus the posterior mean is (x + 0.5)/(n + 1).
The default type of interval constructed is "highest" which computes the highest probability density (hpd) interval which assures the shortest interval possible. The hpd intervals will achieve a probability that is within tol of the specified conf.level. Setting type to "central" constructs intervals that have equal tail probabilities.
If 0 or n successes are observed, a one-sided confidence interval is returned.
Value
For binom.bayes, a data.frame containing the observed
proportions and the lower and upper bounds of the confidence interval.
For binom.bayes.densityplot, a ggplot object that can
printed to a graphics device, or have additional layers added.
References
Gelman, A., Carlin, J. B., Stern, H. S., and Rubin, D. B. (1997) Bayesian Data Analysis, London, U.K.: Chapman and Hall.
See also
Examples
# Example using highest probability density.
hpd <- binom.bayes(
x = 0:10, n = 10, type = "highest", conf.level = 0.8, tol = 1e-9)
print(hpd)
#> method x n shape1 shape2 mean lower upper sig
#> 1 bayes 0 10 0.5 10.5 0.04545455 0.000000000 0.0769566 0.2
#> 2 bayes 1 10 1.5 9.5 0.13636364 0.003771457 0.2145408 0.2
#> 3 bayes 2 10 2.5 8.5 0.22727273 0.053633312 0.3494770 0.2
#> 4 bayes 3 10 3.5 7.5 0.31818182 0.127646004 0.4728042 0.2
#> 5 bayes 4 10 4.5 6.5 0.40909091 0.214091147 0.5858428 0.2
#> 6 bayes 5 10 5.5 5.5 0.50000000 0.309892028 0.6901080 0.2
#> 7 bayes 6 10 6.5 4.5 0.59090909 0.414157185 0.7859089 0.2
#> 8 bayes 7 10 7.5 3.5 0.68181818 0.527195809 0.8723540 0.2
#> 9 bayes 8 10 8.5 2.5 0.77272727 0.650522982 0.9463667 0.2
#> 10 bayes 9 10 9.5 1.5 0.86363636 0.785459228 0.9962285 0.2
#> 11 bayes 10 10 10.5 0.5 0.95454545 0.923043403 1.0000000 0.2
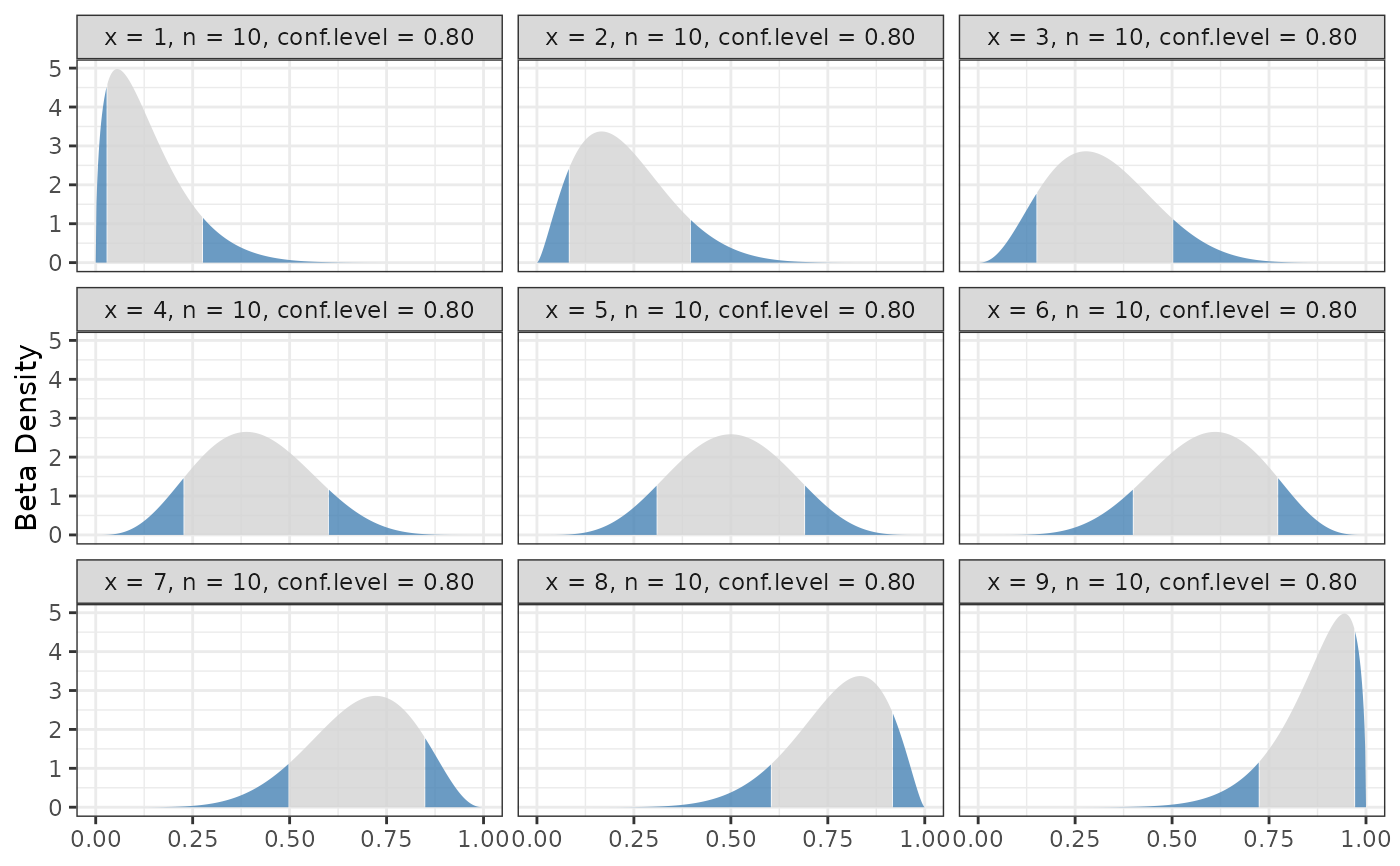
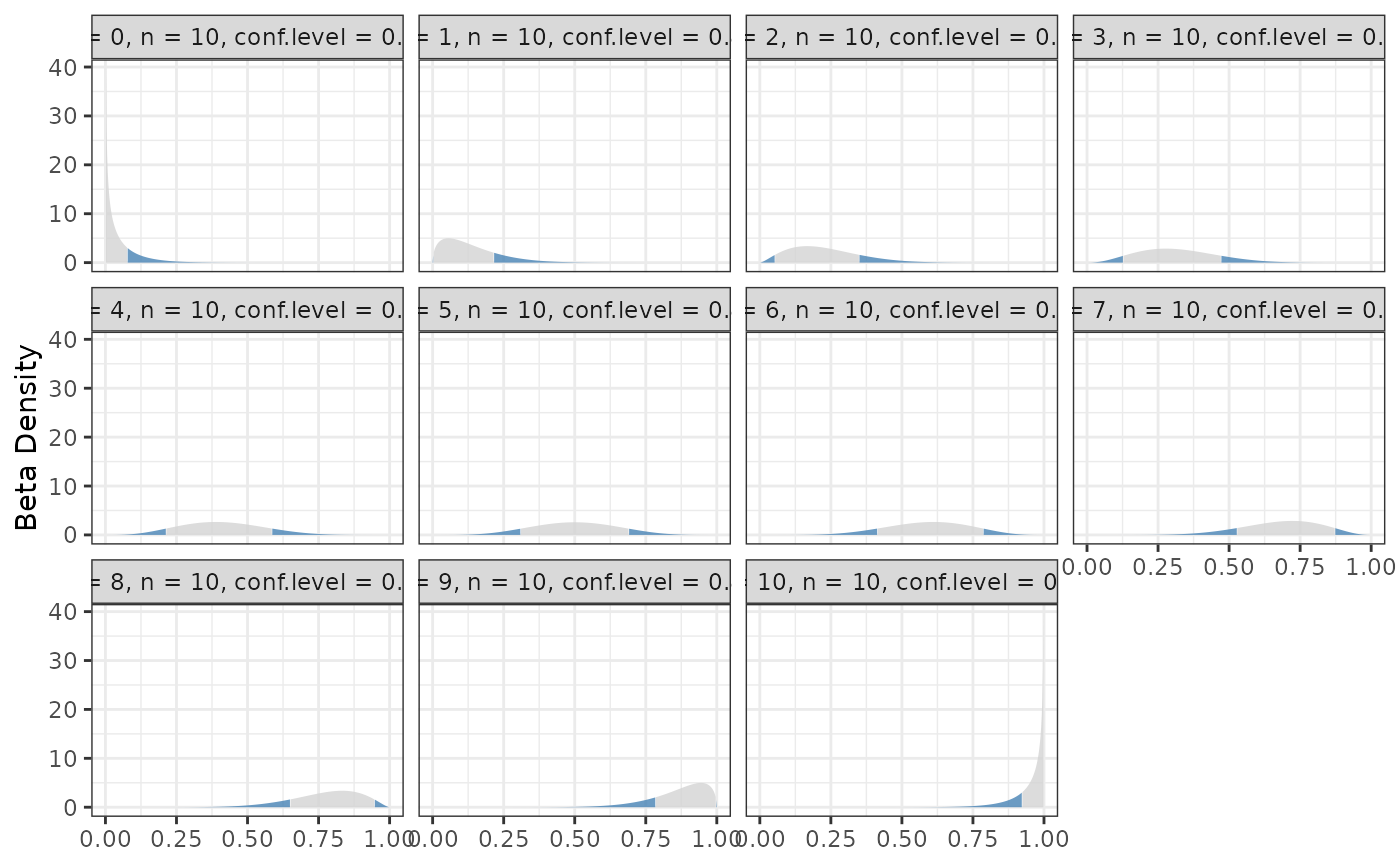
binom.bayes.densityplot(hpd)
#> Loading required namespace: ggplot2
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the binom package.
#> Please report the issue to the authors.
 # Remove the extremes from the plot since they make things hard
# to see.
binom.bayes.densityplot(hpd[hpd$x != 0 & hpd$x != 10, ])
# Remove the extremes from the plot since they make things hard
# to see.
binom.bayes.densityplot(hpd[hpd$x != 0 & hpd$x != 10, ])
 # Example using central probability.
central <- binom.bayes(
x = 0:10, n = 10, type = "central", conf.level = 0.8, tol = 1e-9)
print(central)
#> method x n shape1 shape2 mean lower upper sig
#> 1 bayes 0 10 0.5 10.5 0.04545455 0.00000000 0.0769566 0.2
#> 2 bayes 1 10 1.5 9.5 0.13636364 0.02954113 0.2745618 0.2
#> 3 bayes 2 10 2.5 8.5 0.22727273 0.08361516 0.3948296 0.2
#> 4 bayes 3 10 3.5 7.5 0.31818182 0.15059123 0.5017811 0.2
#> 5 bayes 4 10 4.5 6.5 0.40909091 0.22651679 0.5996844 0.2
#> 6 bayes 5 10 5.5 5.5 0.50000000 0.30989203 0.6901080 0.2
#> 7 bayes 6 10 6.5 4.5 0.59090909 0.40031562 0.7734832 0.2
#> 8 bayes 7 10 7.5 3.5 0.68181818 0.49821888 0.8494088 0.2
#> 9 bayes 8 10 8.5 2.5 0.77272727 0.60517036 0.9163848 0.2
#> 10 bayes 9 10 9.5 1.5 0.86363636 0.72543825 0.9704589 0.2
#> 11 bayes 10 10 10.5 0.5 0.95454545 0.92304340 1.0000000 0.2
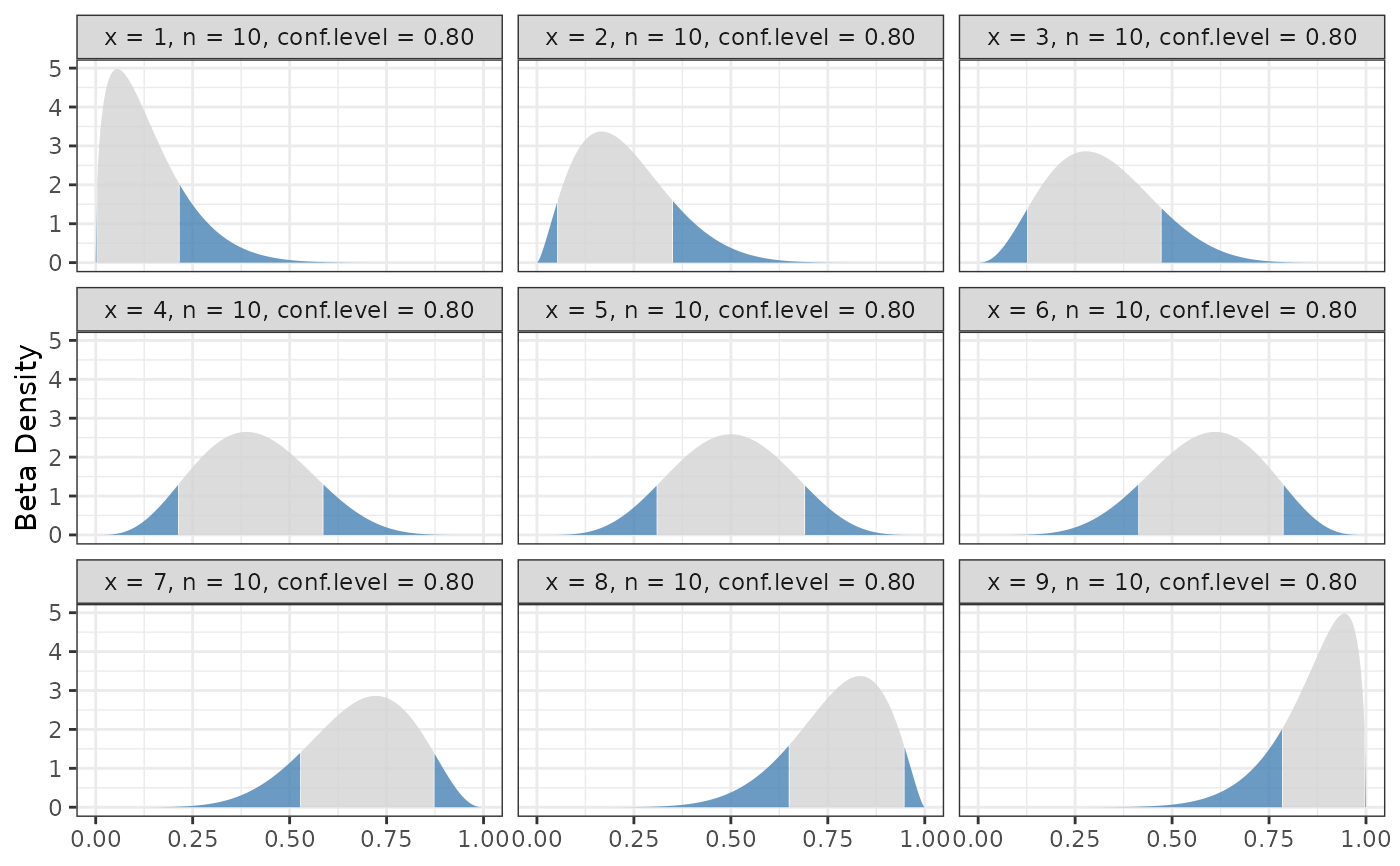
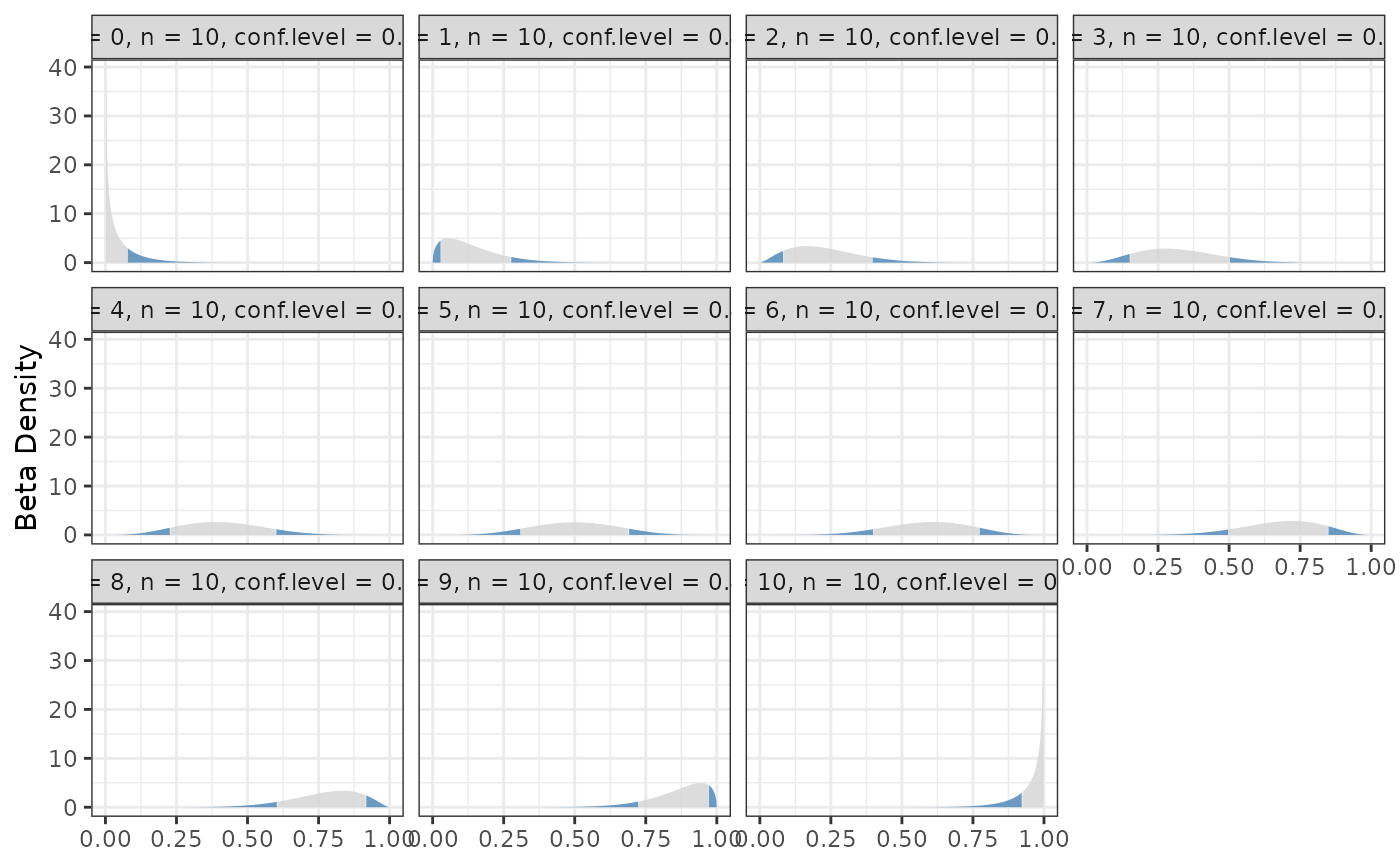
binom.bayes.densityplot(central)
# Example using central probability.
central <- binom.bayes(
x = 0:10, n = 10, type = "central", conf.level = 0.8, tol = 1e-9)
print(central)
#> method x n shape1 shape2 mean lower upper sig
#> 1 bayes 0 10 0.5 10.5 0.04545455 0.00000000 0.0769566 0.2
#> 2 bayes 1 10 1.5 9.5 0.13636364 0.02954113 0.2745618 0.2
#> 3 bayes 2 10 2.5 8.5 0.22727273 0.08361516 0.3948296 0.2
#> 4 bayes 3 10 3.5 7.5 0.31818182 0.15059123 0.5017811 0.2
#> 5 bayes 4 10 4.5 6.5 0.40909091 0.22651679 0.5996844 0.2
#> 6 bayes 5 10 5.5 5.5 0.50000000 0.30989203 0.6901080 0.2
#> 7 bayes 6 10 6.5 4.5 0.59090909 0.40031562 0.7734832 0.2
#> 8 bayes 7 10 7.5 3.5 0.68181818 0.49821888 0.8494088 0.2
#> 9 bayes 8 10 8.5 2.5 0.77272727 0.60517036 0.9163848 0.2
#> 10 bayes 9 10 9.5 1.5 0.86363636 0.72543825 0.9704589 0.2
#> 11 bayes 10 10 10.5 0.5 0.95454545 0.92304340 1.0000000 0.2
binom.bayes.densityplot(central)
 # Remove the extremes from the plot since they make things hard
# to see.
binom.bayes.densityplot(central[central$x != 0 & central$x != 10, ])
# Remove the extremes from the plot since they make things hard
# to see.
binom.bayes.densityplot(central[central$x != 0 & central$x != 10, ])