Trivariate Normal Distribution Density and Random Variates
trinormalUC.RdDensity and random generation for the trivariate normal distribution distribution.
Usage
dtrinorm(x1, x2, x3, mean1 = 0, mean2 = 0, mean3 = 0,
var1 = 1, var2 = 1, var3 = 1,
cov12 = 0, cov23 = 0, cov13 = 0, log = FALSE)
rtrinorm(n, mean1 = 0, mean2 = 0, mean3 = 0,
var1 = 1, var2 = 1, var3 = 1,
cov12 = 0, cov23 = 0, cov13 = 0)Arguments
- x1, x2, x3
vector of quantiles.
- mean1, mean2, mean3
vectors of means.
- var1, var2, var3
vectors of variances.
- cov12, cov23, cov13
vectors of covariances.
- n
number of observations. Same as
rnorm.- log
Logical. If
log = TRUEthen the logarithm of the density is returned.
Details
The default arguments correspond to the standard trivariate normal
distribution with correlation parameters equal to 0,
which corresponds to three independent standard normal distributions.
Let sd1 (say) be sqrt(var1) and
written \(\sigma_1\), etc.
Then the general formula for each correlation coefficient is
of the form
\(\rho_{12} = cov_{12} / (\sigma_1 \sigma_2)\),
and similarly for the two others.
Thus if the var arguments are left alone then
the cov can be inputted with \(\rho\)s.
Warning
dtrinorm()'s arguments might change in the future!
It's safest to use the full argument names
to future-proof possible changes!
Note
For rtrinorm(),
if the \(i\)th variance-covariance matrix is not
positive-definite then the \(i\)th row is all NAs.
Examples
if (FALSE) nn <- 1000
tdata <- data.frame(x2 = sort(runif(nn)))
tdata <- transform(tdata, mean1 = 1 + 2 * x2,
mean2 = 3 + 1 * x2, mean3 = 4,
var1 = exp( 1), var2 = exp( 1), var3 = exp( 1),
rho12 = rhobitlink( 1, inverse = TRUE),
rho23 = rhobitlink( 1, inverse = TRUE),
rho13 = rhobitlink(-1, inverse = TRUE))
ymat <- with(tdata, rtrinorm(nn, mean1, mean2, mean3,
var1, var2, var3,
sqrt(var1)*sqrt(var1)*rho12,
sqrt(var2)*sqrt(var3)*rho23,
sqrt(var1)*sqrt(var3)*rho13))
pairs(ymat, col = "blue")
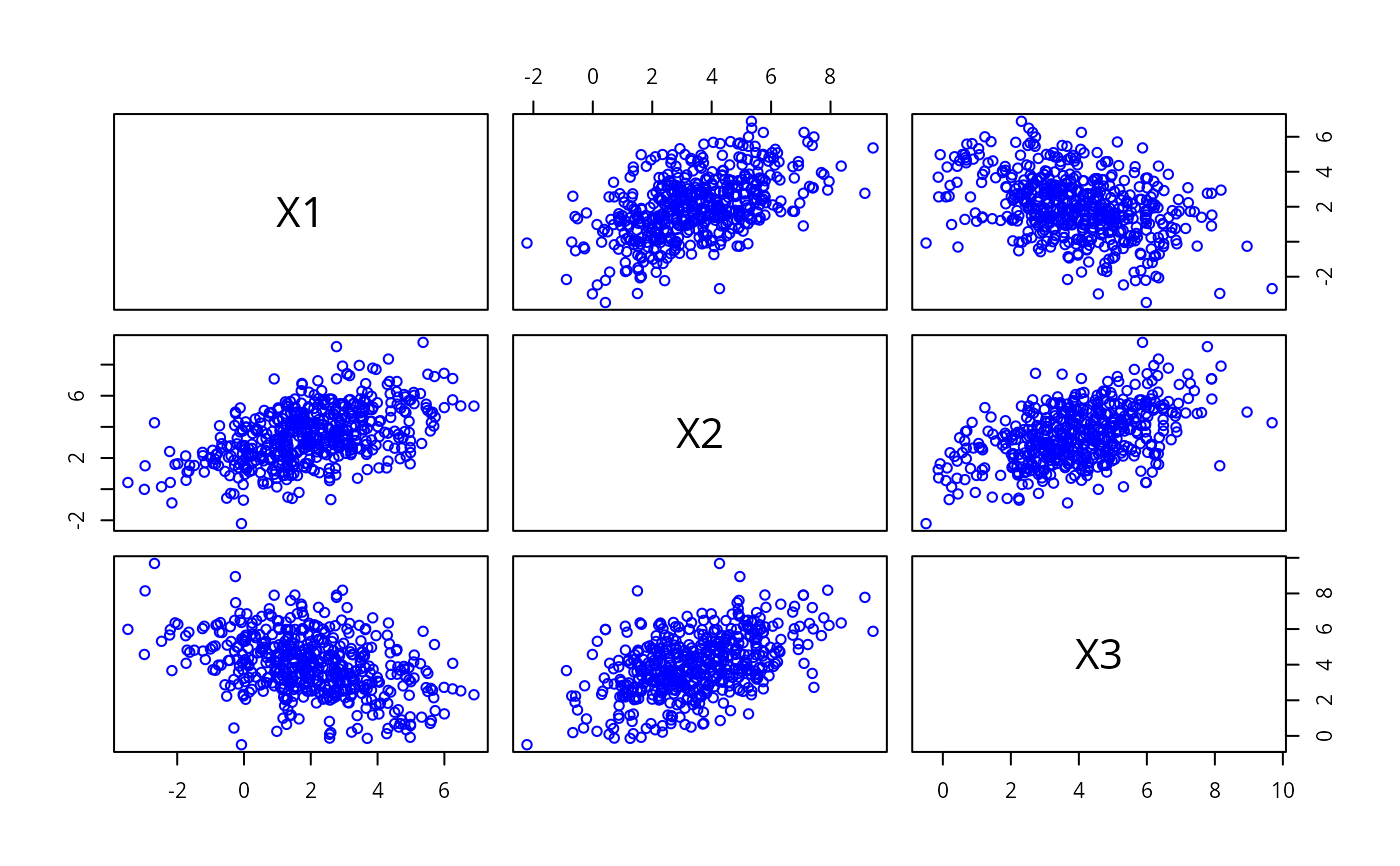 # \dontrun{}
# \dontrun{}