Probit Link Function
probitlink.RdComputes the probit transformation, including its inverse and the first two derivatives.
Details
The probit link function is commonly used for parameters that
lie in the unit interval.
It is the inverse CDF of the standard normal distribution.
Numerical values of theta close to 0 or 1 or out of range
result in
Inf, -Inf, NA or NaN.
Value
For deriv = 0, the probit of theta, i.e.,
qnorm(theta) when inverse = FALSE, and if inverse =
TRUE then pnorm(theta).
For deriv = 1, then the function returns
d eta / d theta as a function of theta
if inverse = FALSE,
else if inverse = TRUE then it returns the reciprocal.
References
McCullagh, P. and Nelder, J. A. (1989). Generalized Linear Models, 2nd ed. London: Chapman & Hall.
Note
Numerical instability may occur when theta is close to 1 or 0.
One way of overcoming this is to use bvalue.
In terms of the threshold approach with cumulative probabilities for
an ordinal response this link function corresponds to the univariate
normal distribution (see uninormal).
Examples
p <- seq(0.01, 0.99, by = 0.01)
probitlink(p)
#> [1] -2.32634787 -2.05374891 -1.88079361 -1.75068607 -1.64485363 -1.55477359
#> [7] -1.47579103 -1.40507156 -1.34075503 -1.28155157 -1.22652812 -1.17498679
#> [13] -1.12639113 -1.08031934 -1.03643339 -0.99445788 -0.95416525 -0.91536509
#> [19] -0.87789630 -0.84162123 -0.80642125 -0.77219321 -0.73884685 -0.70630256
#> [25] -0.67448975 -0.64334541 -0.61281299 -0.58284151 -0.55338472 -0.52440051
#> [31] -0.49585035 -0.46769880 -0.43991317 -0.41246313 -0.38532047 -0.35845879
#> [37] -0.33185335 -0.30548079 -0.27931903 -0.25334710 -0.22754498 -0.20189348
#> [43] -0.17637416 -0.15096922 -0.12566135 -0.10043372 -0.07526986 -0.05015358
#> [49] -0.02506891 0.00000000 0.02506891 0.05015358 0.07526986 0.10043372
#> [55] 0.12566135 0.15096922 0.17637416 0.20189348 0.22754498 0.25334710
#> [61] 0.27931903 0.30548079 0.33185335 0.35845879 0.38532047 0.41246313
#> [67] 0.43991317 0.46769880 0.49585035 0.52440051 0.55338472 0.58284151
#> [73] 0.61281299 0.64334541 0.67448975 0.70630256 0.73884685 0.77219321
#> [79] 0.80642125 0.84162123 0.87789630 0.91536509 0.95416525 0.99445788
#> [85] 1.03643339 1.08031934 1.12639113 1.17498679 1.22652812 1.28155157
#> [91] 1.34075503 1.40507156 1.47579103 1.55477359 1.64485363 1.75068607
#> [97] 1.88079361 2.05374891 2.32634787
max(abs(probitlink(probitlink(p), inverse = TRUE) - p)) # Should be 0
#> [1] 1.110223e-16
p <- c(seq(-0.02, 0.02, by = 0.01), seq(0.97, 1.02, by = 0.01))
probitlink(p) # Has NAs
#> [1] NaN NaN -Inf -2.326348 -2.053749 1.880794 2.053749
#> [8] 2.326348 Inf NaN NaN
probitlink(p, bvalue = .Machine$double.eps) # Has no NAs
#> [1] -8.125891 -8.125891 -8.125891 -2.326348 -2.053749 1.880794 2.053749
#> [8] 2.326348 8.125891 8.125891 8.125891
if (FALSE) p <- seq(0.01, 0.99, by = 0.01); par(lwd = (mylwd <- 2))
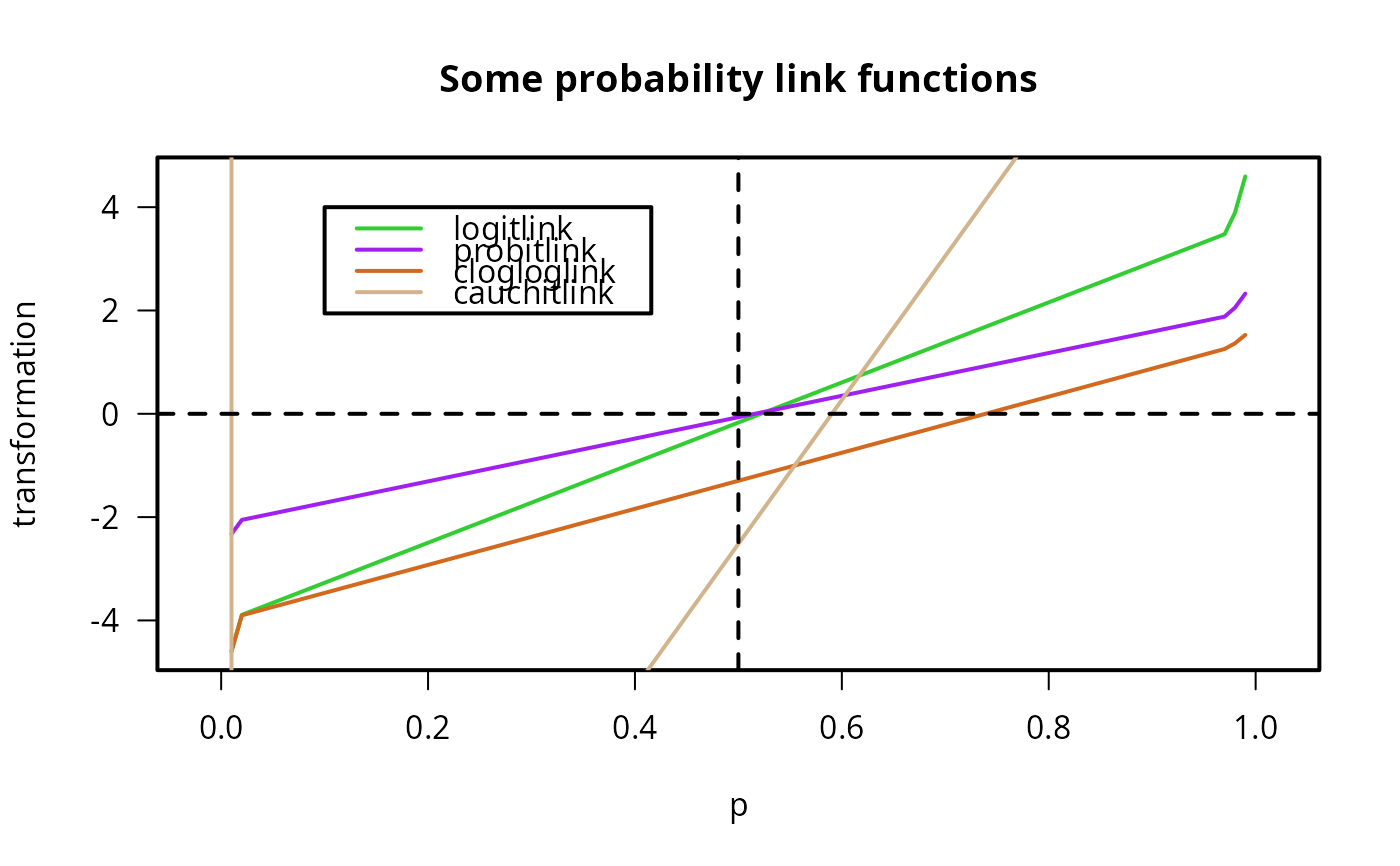
plot(p, logitlink(p), type = "l", col = "limegreen", ylab = "transformation",
las = 1, main = "Some probability link functions")
lines(p, probitlink(p), col = "purple")
lines(p, clogloglink(p), col = "chocolate")
lines(p, cauchitlink(p), col = "tan")
abline(v = 0.5, h = 0, lty = "dashed")
legend(0.1, 4, c("logitlink", "probitlink", "clogloglink", "cauchitlink"),
col = c("limegreen", "purple", "chocolate", "tan"), lwd = mylwd)
 par(lwd = 1) # \dontrun{}
par(lwd = 1) # \dontrun{}