Nakagami Regression Family Function
nakagami.RdEstimation of the two parameters of the Nakagami distribution by maximum likelihood estimation.
Usage
nakagami(lscale = "loglink", lshape = "loglink", iscale = 1,
ishape = NULL, nowarning = FALSE, zero = "shape")Arguments
- nowarning
Logical. Suppress a warning?
- lscale, lshape
Parameter link functions applied to the scale and shape parameters. Log links ensure they are positive. See
Linksfor more choices and information.- iscale, ishape
Optional initial values for the shape and scale parameters. For
ishape, aNULLvalue means it is obtained in theinitializeslot based on the value ofiscale. Foriscale, assigning aNULLmeans a value is obtained in theinitializeslot, however, setting another numerical value is recommended if convergence fails or is too slow.- zero
Details
The Nakagami distribution, which is useful for modelling wireless systems such as radio links, can be written $$f(y) = 2 (shape/scale)^{shape} y^{2 \times shape-1} \exp(-shape \times y^2/scale) / \Gamma(shape)$$ for \(y > 0\), \(shape > 0\), \(scale > 0\). The mean of \(Y\) is \(\sqrt{scale/shape} \times \Gamma(shape+0.5) / \Gamma(shape)\) and these are returned as the fitted values. By default, the linear/additive predictors are \(\eta_1=\log(scale)\) and \(\eta_2=\log(shape)\). Fisher scoring is implemented.
Value
An object of class "vglmff" (see vglmff-class).
The object is used by modelling functions such as vglm,
and vgam.
References
Nakagami, M. (1960). The m-distribution: a general formula of intensity distribution of rapid fading, pp.3–36 in: Statistical Methods in Radio Wave Propagation. W. C. Hoffman, Ed., New York: Pergamon.
Note
The Nakagami distribution is also known as the Nakagami-m distribution, where \(m=shape\) here. Special cases: \(m=0.5\) is a one-sided Gaussian distribution and \(m=1\) is a Rayleigh distribution. The second moment is \(E(Y^2)=m\).
If \(Y\) has a Nakagami distribution with parameters shape and scale then \(Y^2\) has a gamma distribution with shape parameter shape and scale parameter scale/shape.
Examples
nn <- 1000; shape <- exp(0); Scale <- exp(1)
ndata <- data.frame(y1 = sqrt(rgamma(nn, shape = shape, scale = Scale/shape)))
nfit <- vglm(y1 ~ 1, nakagami, data = ndata, trace = TRUE, crit = "coef")
#> Iteration 1: coefficients = -0.29687619, -2.15881032
#> Iteration 2: coefficients = 2.3237907, -1.5218807
#> Iteration 3: coefficients = 1.5872099, -0.9661463
#> Iteration 4: coefficients = 1.13743497, -0.46826416
#> Iteration 5: coefficients = 1.00016546, -0.12708838
#> Iteration 6: coefficients = 0.989835766, -0.022320279
#> Iteration 7: coefficients = 0.98978205, -0.01519528
#> Iteration 8: coefficients = 0.989782044, -0.015165205
#> Iteration 9: coefficients = 0.989782044, -0.015165205
ndata <- transform(ndata, y2 = rnaka(nn, scale = Scale, shape = shape))
nfit <- vglm(y2 ~ 1, nakagami(iscale = 3), data = ndata, trace = TRUE)
#> Iteration 1: loglikelihood = -8946.7171
#> Iteration 2: loglikelihood = -7948.6648
#> Iteration 3: loglikelihood = -6953.3186
#> Iteration 4: loglikelihood = -5964.2835
#> Iteration 5: loglikelihood = -4989.4468
#> Iteration 6: loglikelihood = -4045.0096
#> Iteration 7: loglikelihood = -3160.8638
#> Iteration 8: loglikelihood = -2383.297
#> Iteration 9: loglikelihood = -1767.0711
#> Iteration 10: loglikelihood = -1356.6308
#> Iteration 11: loglikelihood = -1163.1982
#> Iteration 12: loglikelihood = -1121.0429
#> Iteration 13: loglikelihood = -1119.1599
#> Iteration 14: loglikelihood = -1119.1561
#> Iteration 15: loglikelihood = -1119.1561
head(fitted(nfit))
#> [,1]
#> [1,] 1.494389
#> [2,] 1.494389
#> [3,] 1.494389
#> [4,] 1.494389
#> [5,] 1.494389
#> [6,] 1.494389
with(ndata, mean(y2))
#> [1] 1.495657
coef(nfit, matrix = TRUE)
#> loglink(scale) loglink(shape)
#> (Intercept) 1.045623 -0.002743848
(Cfit <- Coef(nfit))
#> scale shape
#> 2.8451716 0.9972599
if (FALSE) sy <- with(ndata, sort(y2))
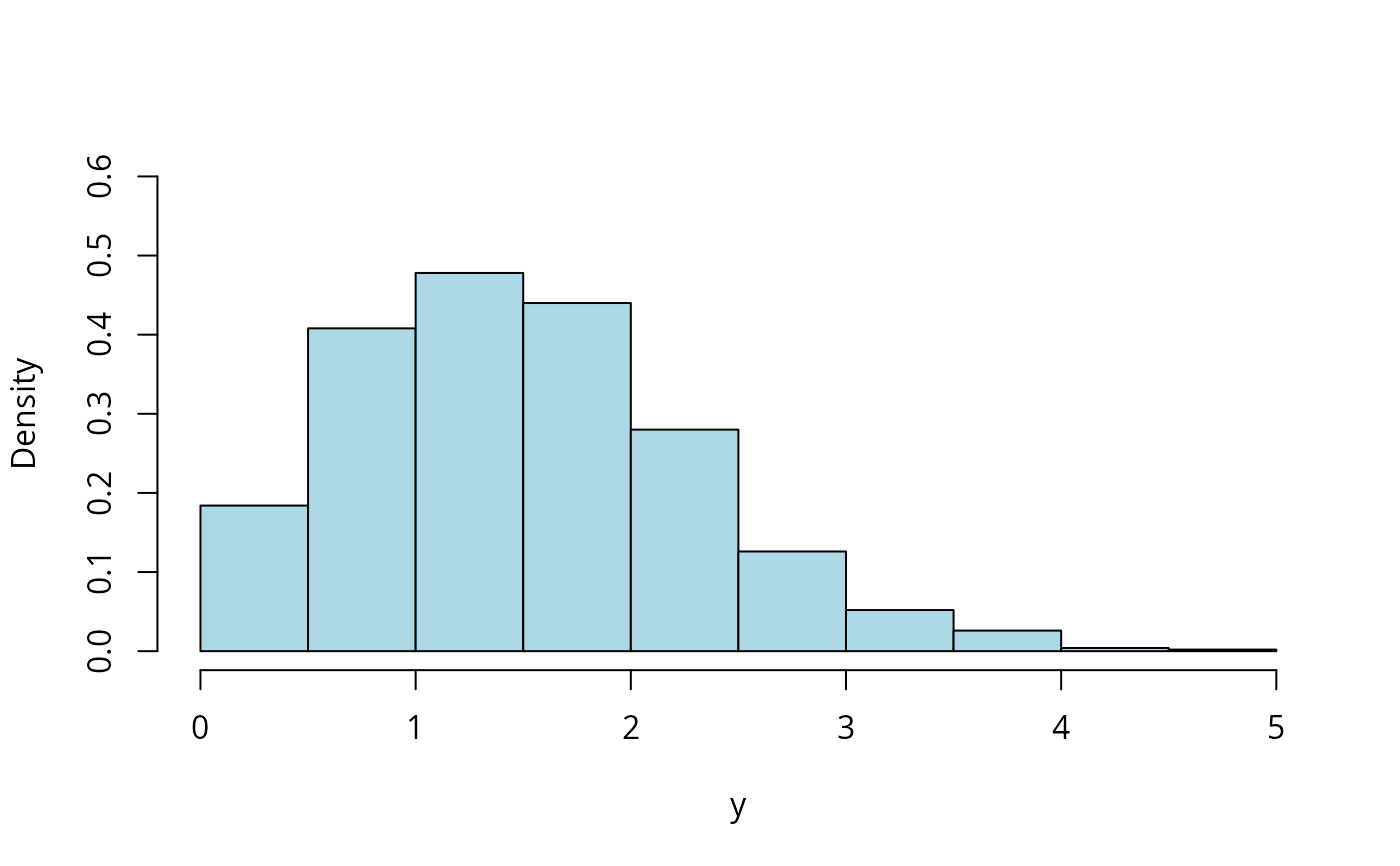
hist(with(ndata, y2), prob = TRUE, main = "", xlab = "y", ylim = c(0, 0.6),
col = "lightblue")
 lines(dnaka(sy, scale = Cfit["scale"], shape = Cfit["shape"]) ~ sy,
data = ndata, col = "orange") # \dontrun{}
#> Error in eval(predvars, data, env): object 'sy' not found
lines(dnaka(sy, scale = Cfit["scale"], shape = Cfit["shape"]) ~ sy,
data = ndata, col = "orange") # \dontrun{}
#> Error in eval(predvars, data, env): object 'sy' not found