Doctoral Publications
PhDPublications.RdCross-section data on the scientific productivity of PhD students in biochemistry.
Usage
data("PhDPublications")Format
A data frame containing 915 observations on 6 variables.
- articles
Number of articles published during last 3 years of PhD.
- gender
factor indicating gender.
- married
factor. Is the PhD student married?
- kids
Number of children less than 6 years old.
- prestige
Prestige of the graduate program.
- mentor
Number of articles published by student's mentor.
References
Long, J.S. (1990). Regression Models for Categorical and Limited Dependent Variables. Thousand Oaks: Sage Publications.
Long, J.S. (1997). The Origin of Sex Differences in Science. Social Forces, 68, 1297–1315.
Examples
## from Long (1997)
data("PhDPublications")
## Table 8.1, p. 227
summary(PhDPublications)
#> articles gender married kids prestige
#> Min. : 0.000 male :494 no :309 Min. :0.0000 Min. :0.755
#> 1st Qu.: 0.000 female:421 yes:606 1st Qu.:0.0000 1st Qu.:2.260
#> Median : 1.000 Median :0.0000 Median :3.150
#> Mean : 1.693 Mean :0.4951 Mean :3.103
#> 3rd Qu.: 2.000 3rd Qu.:1.0000 3rd Qu.:3.920
#> Max. :19.000 Max. :3.0000 Max. :4.620
#> mentor
#> Min. : 0.000
#> 1st Qu.: 3.000
#> Median : 6.000
#> Mean : 8.767
#> 3rd Qu.:12.000
#> Max. :77.000
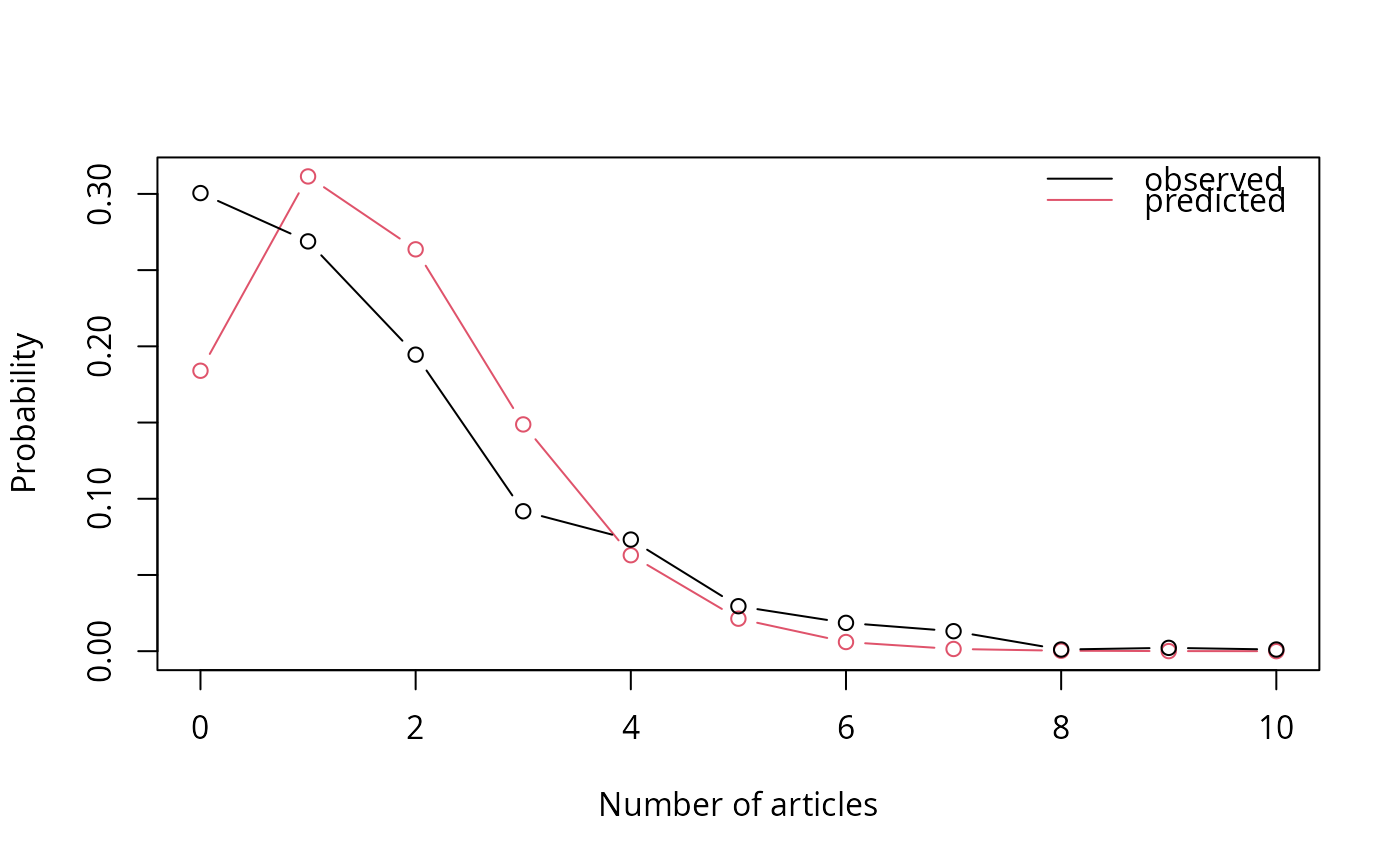
## Figure 8.2, p. 220
plot(0:10, dpois(0:10, mean(PhDPublications$articles)), type = "b", col = 2,
xlab = "Number of articles", ylab = "Probability")
lines(0:10, prop.table(table(PhDPublications$articles))[1:11], type = "b")
legend("topright", c("observed", "predicted"), col = 1:2, lty = rep(1, 2), bty = "n")
 ## Table 8.2, p. 228
fm_lrm <- lm(log(articles + 0.5) ~ ., data = PhDPublications)
summary(fm_lrm)
#>
#> Call:
#> lm(formula = log(articles + 0.5) ~ ., data = PhDPublications)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.87006 -0.87012 0.07973 0.63630 2.17374
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.177774 0.107725 1.650 0.0992 .
#> genderfemale -0.134567 0.057298 -2.349 0.0191 *
#> marriedyes 0.132826 0.065027 2.043 0.0414 *
#> kids -0.133148 0.040655 -3.275 0.0011 **
#> prestige 0.025502 0.028469 0.896 0.3706
#> mentor 0.025421 0.002954 8.607 <2e-16 ***
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#>
#> Residual standard error: 0.8146 on 909 degrees of freedom
#> Multiple R-squared: 0.1008, Adjusted R-squared: 0.09582
#> F-statistic: 20.37 on 5 and 909 DF, p-value: < 2.2e-16
#>
-2 * logLik(fm_lrm)
#> 'log Lik.' 2215.323 (df=7)
fm_prm <- glm(articles ~ ., data = PhDPublications, family = poisson)
library("MASS")
fm_nbrm <- glm.nb(articles ~ ., data = PhDPublications)
## Table 8.3, p. 246
library("pscl")
fm_zip <- zeroinfl(articles ~ . | ., data = PhDPublications)
fm_zinb <- zeroinfl(articles ~ . | ., data = PhDPublications, dist = "negbin")
## Table 8.2, p. 228
fm_lrm <- lm(log(articles + 0.5) ~ ., data = PhDPublications)
summary(fm_lrm)
#>
#> Call:
#> lm(formula = log(articles + 0.5) ~ ., data = PhDPublications)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.87006 -0.87012 0.07973 0.63630 2.17374
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 0.177774 0.107725 1.650 0.0992 .
#> genderfemale -0.134567 0.057298 -2.349 0.0191 *
#> marriedyes 0.132826 0.065027 2.043 0.0414 *
#> kids -0.133148 0.040655 -3.275 0.0011 **
#> prestige 0.025502 0.028469 0.896 0.3706
#> mentor 0.025421 0.002954 8.607 <2e-16 ***
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#>
#> Residual standard error: 0.8146 on 909 degrees of freedom
#> Multiple R-squared: 0.1008, Adjusted R-squared: 0.09582
#> F-statistic: 20.37 on 5 and 909 DF, p-value: < 2.2e-16
#>
-2 * logLik(fm_lrm)
#> 'log Lik.' 2215.323 (df=7)
fm_prm <- glm(articles ~ ., data = PhDPublications, family = poisson)
library("MASS")
fm_nbrm <- glm.nb(articles ~ ., data = PhDPublications)
## Table 8.3, p. 246
library("pscl")
fm_zip <- zeroinfl(articles ~ . | ., data = PhDPublications)
fm_zinb <- zeroinfl(articles ~ . | ., data = PhDPublications, dist = "negbin")